Beyond the Test Tube: Designing Electrochemical Aptamer-Based (EAB) Sensors That Actually Work
If you're working with Electrochemical Aptamer-Based (EAB) sensors, you know the power of aptamers—those single-stranded DNA or RNA molecules that bind to a specific target. But there's a catch: the classic method for finding these aptamers isn't perfectly suited for the unique demands of electrochemical sensing.
In this post, we’ll break down the gap between traditional aptamer selection and the real-world needs of EAB sensors, and outline a practical path to identifying the best aptamer for your electrochemical assay.
A Quick Refresher: How Do We Get Aptamers?
The standard process is called SELEX (Systematic Evolution of Ligands by EXponential enrichment). Here’s how it works in broad strokes:
Library: You start with a vast, random pool of single-stranded DNA or RNA sequences.
Binding: You introduce your target molecule (analyte), and a small fraction of these sequences will bind to it.
Partitioning: You separate the sequences that bound successfully from those that didn't.
Amplification: You use PCR (for DNA) or RT-PCR (for RNA) to make millions of copies of the binding sequences.
Iteration: You repeat this cycle several times, each time enriching the pool with sequences that have the strongest affinity for your target.
SELEX is brilliant at one thing: optimizing for high-affinity binding in a solution (homogeneous phase).
The Problem: Your Sensor Isn't a Test Tube
This is where the challenge for EAB development begins. In reality, your electrochemical sensor operates in a heterogeneous phase:
The aptamer is tethered to a surface (like an electrode).
The binding event happens at this interface.
You need this binding to produce a measurable electrochemical signal.
SELEX, in its classic form, doesn't account for any of this. Here’s why that matters:
The Surface Environment is Different: The local pH, salt concentration, and the electrical "double layer" near the electrode can alter the aptamer's structure and function. SELEX wasn't run in these conditions.
Signal Generation is Key: For an electrochemical signal to be strong, the reporting molecule (e.g., Methylene Blue) must be within ~4 nanometers of the electrode surface—a distance known as the Debye length. SELEX doesn't select for binding events that bring a tag this close to a surface.
Conformational Change isn't Guaranteed: Many elegant EAB sensors rely on the aptamer changing its shape upon binding, which alters the electron transfer. SELEX only cares about binding, not about a subsequent shape shift that's useful for electrochemistry.
In short, SELEX gives you a champion binder for solution, not necessarily a champion signal-generator for your sensor.
A Practical Path to a Functional EAB Sensor
So, how do you find or choose an aptamer that will work in an EAB format? Here is a practical approach:
1. Start with a Deep Literature Search The good news is there's a wealth of published research on EAB sensors. Don't just look for aptamers for your target; search specifically for "electrochemical biosensors" for it. This will immediately point you toward sequences that have already demonstrated success on a surface.
2. Perform In-Silico Triaging Before you order oligonucleotides, do some computer modeling. This can help you:
Predict Flexibility: Aptamers with inherent flexibility are more likely to undergo a useful conformational change upon binding.
Check for Modify-ability: Can you add a label like Methylene Blue without disrupting the binding pocket? You may need spacers.
Avoid Self-Binding: Look for sequences that might form "secondary structures," causing aptamers to bind to each other on the surface instead of your target.
Model Assay Conditions: Simulate how the aptamer might behave in your actual assay buffer (e.g., interstitial fluid), which may differ from the SELEX buffer.
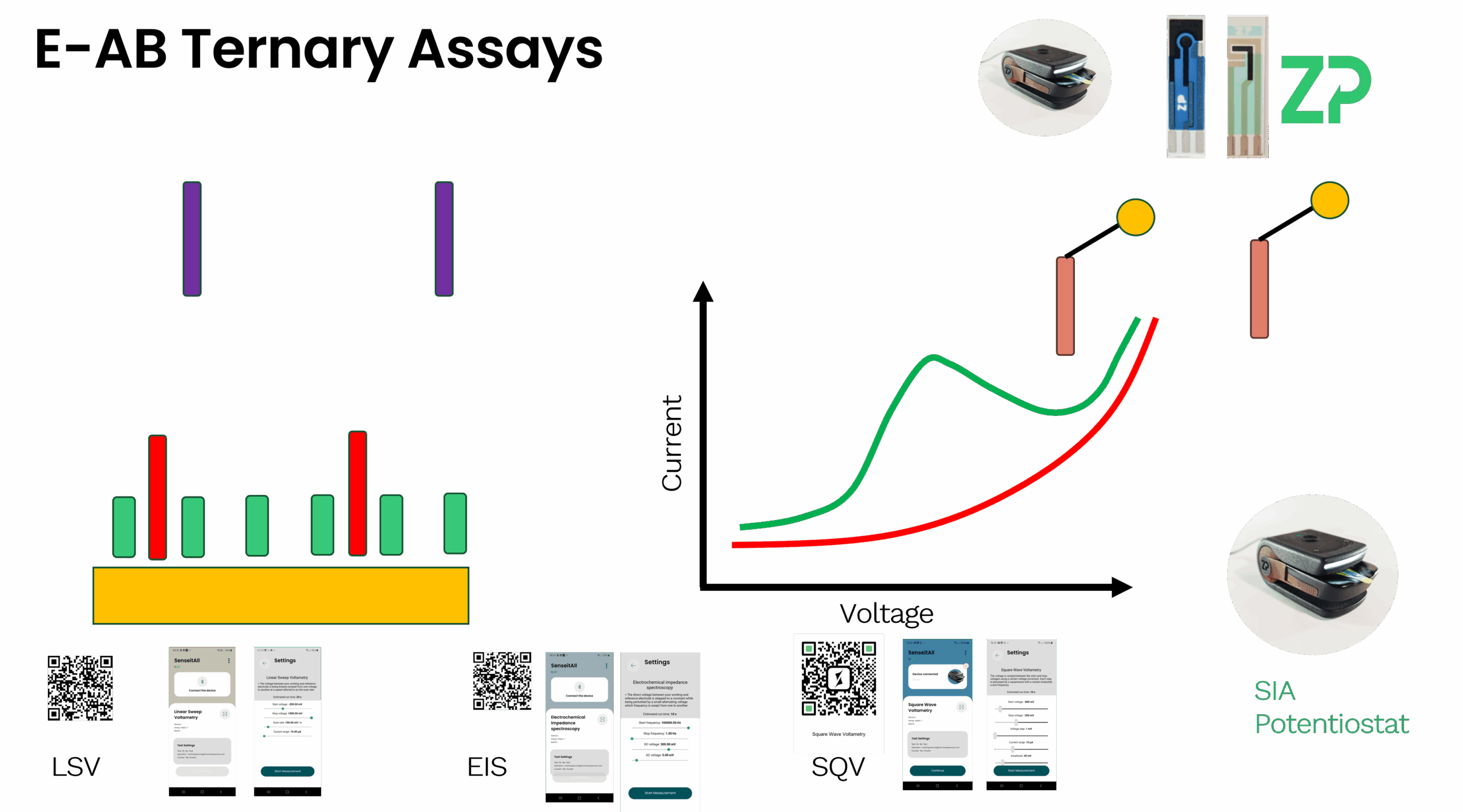
3. Understand Your Signal Mechanism Your choice of aptamer is tied to how you plan to generate a signal. Common EAB strategies include:
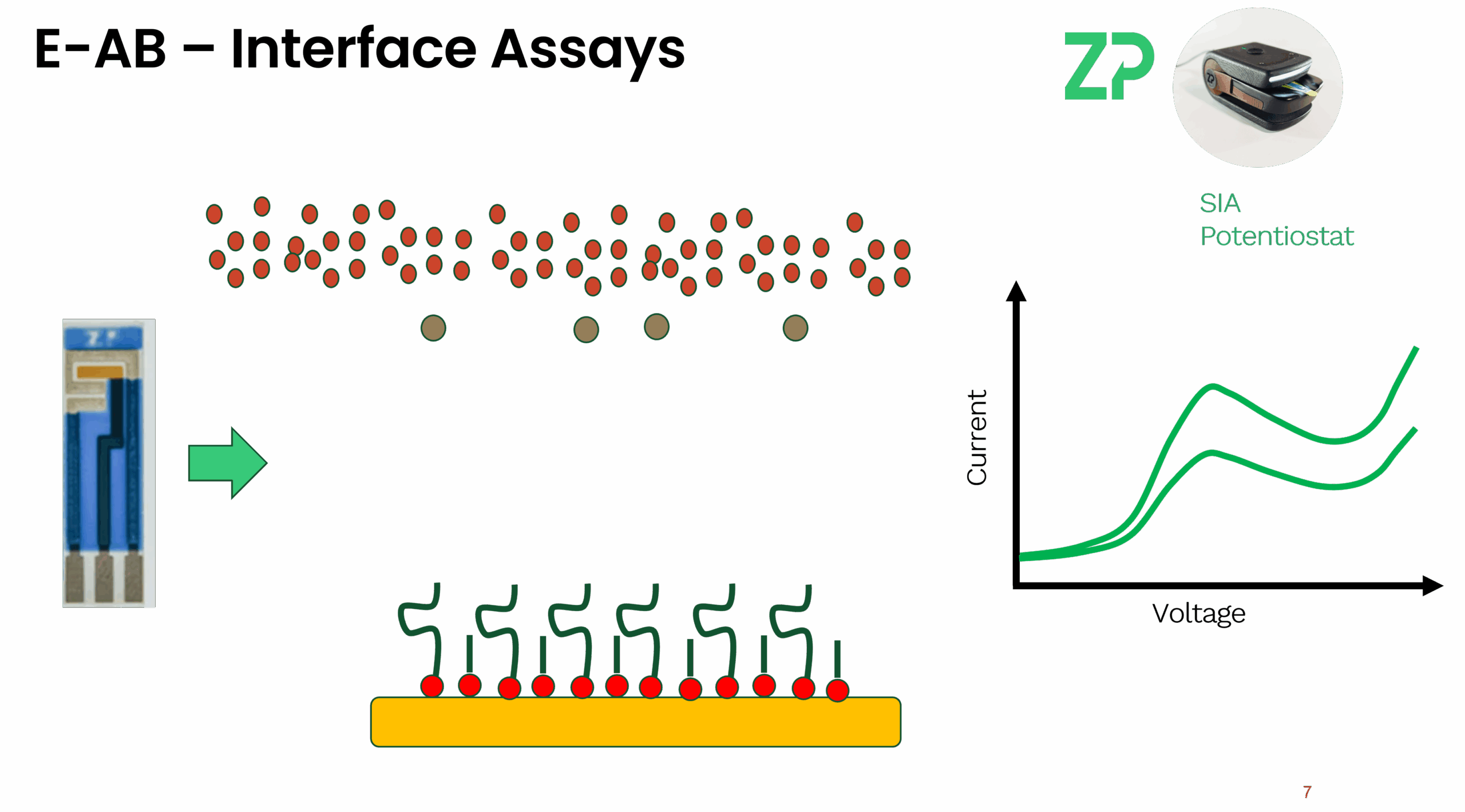
Sandwich Assay: Like an ELISA, you use a capture aptamer on the surface and a reporter aptamer in solution. The target binds both, bringing the reporter tag close to the electrode.
Blocking Assay: The aptamer is tethered to the surface. In its "unbound" state, a redox reporter in solution can easily reach the electrode. Upon target binding, the surface becomes blocked, decreasing the signal.
Conformational Change (The Classic EAB): An aptamer is labeled with a reporter (e.g., Methylene Blue) and designed in a "hairpin." Binding causes a large shape shift that changes the electron transfer efficiency, leading to a measurable change in signal.
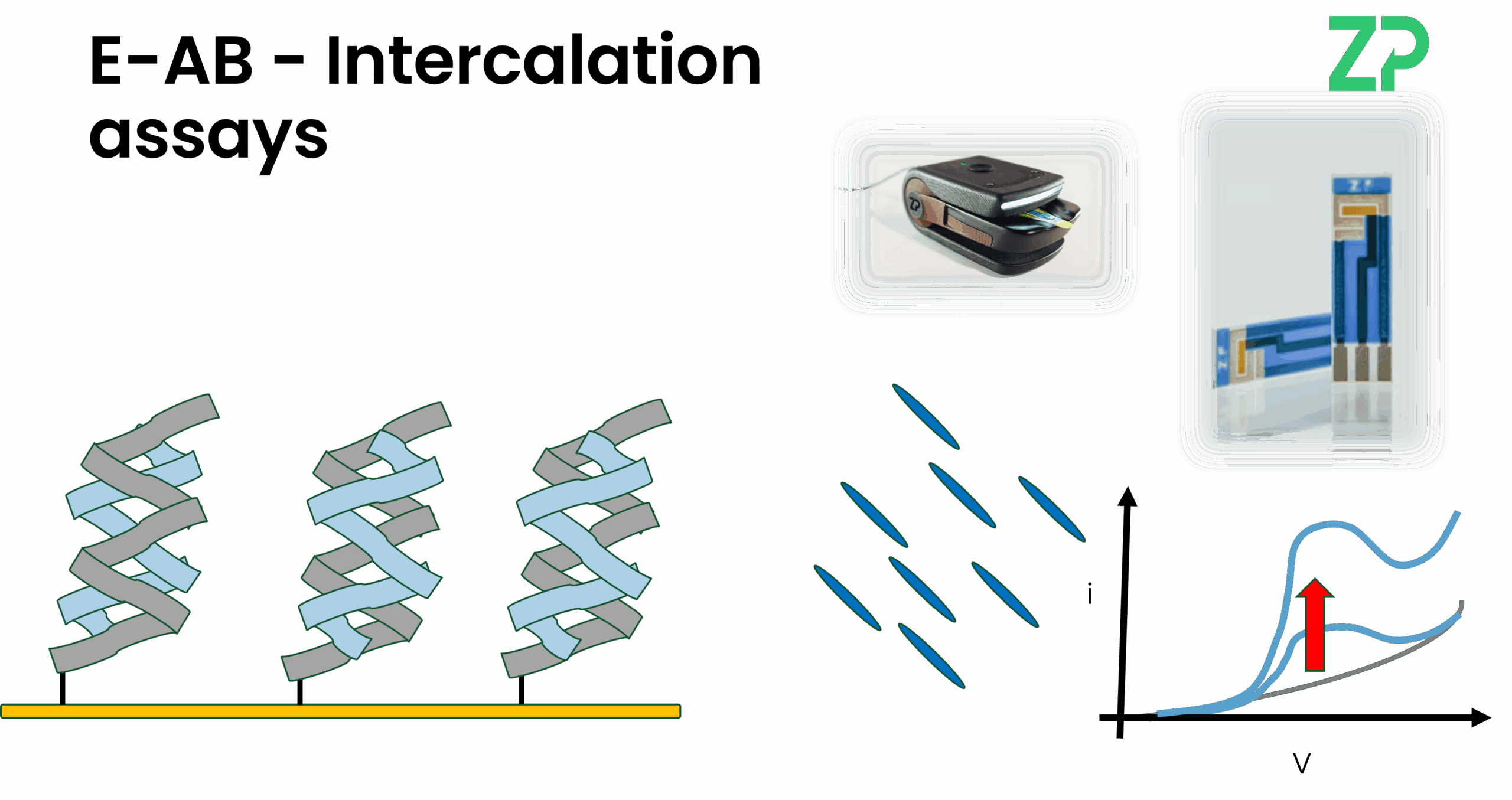
Intercalation: For DNA-based targets, binding can be detected by an intercalating dye (e.g., Methylene Blue) that slips into the DNA duplex, generating a positive signal.
Choosing an aptamer that fits your desired mechanism is crucial.
Key Takeaways
SELEX is a starting point, not the finish line. It optimizes for binding affinity in solution, but an EAB sensor requires optimization for signal generation on a surface.
Leverage the literature to find aptamers that have already shown promise in electrochemical formats.
Use in-silico tools to triage potential aptamers, saving you time and resources in the lab by predicting their behavior in your specific system.
Think about the signal from the start. The best binder is useless if it doesn't produce a clean, measurable electrochemical output.
Developing a robust EAB sensor requires a holistic view that goes beyond the SELEX outcome. By focusing on the practicalities of surface immobilization and signal generation, you can successfully bridge the gap between a promising aptamer and a high-performing sensor.
Ready to start testing your EAB designs? We offer high-quality, research-grade Gold Screen-Printed Electrodes that are perfect for developing and optimizing your electrochemical aptamer-based sensors.